National Action Council for Minorities in Engineering(NACME) Google Applied Machine Learning Intensive (AMLI) at the University of Kentucky
README Developed by:
- Kimi Medina-Castellano -
University of Kentucky
Our task for this project was to create a model to accurately classify samples of microdebitage simulants into 5 tool production stages using numerical data provided by Dr. Phyllis Johnson. The level of desired accuracy was not defined, and this project is currently in development. I tried two models: decision trees and random forest for different Phases and datasets.
Phase 1 indicates the process of using the merged data set of all 5 experiments of chert and obsidian in modeling using decision trees and random forest (multiclass classification).
In this phase, I:
- loaded data: loaded five excel files of measurements of microdebitage samples that represented different stages in the knapping process into separate dataframes.
- checked for missing data: there was no missing data in any of the dataframes.
- assigned an integer for each tool stage: this is the column we will predict on
# add target column, stage, to each dataframe
chert_hh_1['Stage'] = 1
chert_hh_2['Stage'] = 2
chert_sh['Stage'] = 3
obsidian_cr['Stage'] = 4
obsidian_sh['Stage'] = 5
- merged data for tool stages into one large dataframe
# join dataframes
dfs = [chert_hh_1, chert_hh_2, chert_sh, obsidian_cr, obsidian_sh]
tools_df = pd.concat(dfs)
- cleaned the merged dataframe:
# remove unecessary columns + row 0
# row 0 included labels for units of measurement (ex: mm), which were all millimeters
tools_df.drop(columns = ['Id','Filter0','Filter1', 'Filter2', 'Filter3', 'Filter4', 'Filter5', 'Filter6', 'hash'], inplace = True)
tools_df.drop(columns = ['Transparency', 'Curvature', 'Angularity', 'Img Id'], inplace = True)
tools_df.drop(0, inplace = True)
There were three extra columns that our obsidian data had whereas our chert data did not. These were Transparency, Curvature, and Angularity. Initially, I kept these columns for obsidian, added empty columns with the same names to the chert data sets, and filled the empty rows with zeros. Running my two models with this cleaned data gave me 80.1% and 58.7% accuracy in my decision trees and random forest model, respectively. After I tuned the hyperparameters for the decision tree model, my accuracy increased to 80.8%.
We double-checked with Dr. Johnson on whether or not chert microdebitage can be transparent and why there was no data for those three columns. She said they did not have those parameters turned on and should have had that data. This prompted the team to remove the three columns altogether. I ran the updated dataset into both of my models and got about 37% for a basic and tuned model. This caused us to reframe our modeling methods which is discussed in Phase 2. However, let's explore the details behind Phase 1 accuracies below regarding column inclusion and deletion.
Decision trees are supervised machine learning method in which data is split according to certain parameters defined by the user. I used the DecisionTreeClassifier class from sklearn library and had multiple iterations of the trees throughout this process for multiclass classification.
This decision tree model from SciKitLearn was done before I removed the three columns. I first began with a basic model with no tuned parameters that had an accuracy of 80.1%.
# create decision tree
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn import metrics
from sklearn.metrics import classification_report
import matplotlib.pyplot as plt
# set features and target
features = tools_df.loc[:, tools_df.columns != 'Stage'] # included every column except stage
feature_names = features.columns
target = tools_df['Stage']
target_name = 'Stage'
# split data
x_train, x_test, y_train, y_test = train_test_split(
features,
target,
test_size = 0.3,
random_state = 10
)
# train model
decision_tree = DecisionTreeClassifier()
decision_tree.fit(x_train, y_train)
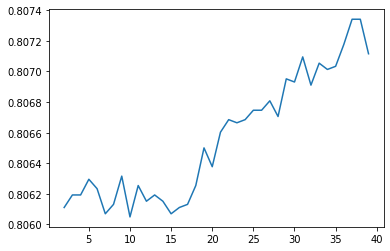
I initially tuned the model by iterating through each integer value a parameter can have within a specific range. For example, here is code of me doing so with the 'min_samples_split' parameter. The result is a graph of the model accuracy vs each integer value of the parameter.
# determine best max_depth
split_list = []
acc_list2 = []
for i in range(2, 40):
dt = tree.DecisionTreeClassifier(
criterion = 'entropy',
max_depth = 11,
min_samples_split = i,
random_state = 10)
# fit tree
dt.fit(x_train, y_train)
# get predictions
y_pred = dt.predict(x_test)
# get accuracy for each iteration and append to list
acc = metrics.accuracy_score(y_test, y_pred)
acc_list2.append(acc)
split_list.append(i)
# show graph of
plt.plot(split_list, acc_list2)
plt.show()
print(max(acc_list2))
for xy in zip(split_list, acc_list2):
print(xy)
I also did this method with the parameters 'max_depth', 'min_samples_leaf', and 'min_weight_fraction_leaf'. I inserted these parameters into a decision tree and got an accuracy of 80.8%. I attempted to use GridSearchCV for decision trees, and it gave a lower accuracy, around 80.7% (although it took a long time to do so). I did not run the random forest model for this scenario because decision trees were more promising.
With the deleted columns, a non-tuned decision tree model had an accuracy of 28.47%. A tuned tree had 36.9% accuracy. We added this line of code which made a big difference:
# drops three columns from merged dataframe
tools_df.drop(columns = ['Transparency', 'Curvature', 'Angularity'], inplace = True)
I decided to test out random forest because it relies on multiple randomly generated decision tree which should, in theory, increase model accuracy and reduce overfitting.
My accuracy for a basic model was 81.98% and a tuned model via GridSearchCV (it worked!) was 81.87%. Below is a basic random forest model:
# create a baseline random forest model
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn import metrics
# determine features and targets
features = tools_df.loc[:, tools_df.columns != 'Stage']
feature_names = features.columns
target = tools_df['Stage']
target_name = 'Stage'
# split data into testing and training data
x_train, x_test, y_train, y_test = train_test_split(
features,
target,
test_size = 0.3,
random_state = 10
)
# create random forest model
rf = RandomForestClassifier()
rf.fit(x_train, y_train)
A basic random forest model without parameter tuning was 37.4% accurate. I was able to use RandomizedSearchCV and get some parameters. The result was a similar accuracy of 37.39%. This didn't feel right, but since we had a better model given our time constraints, I did not explore further. Using RandomizedSearchCV, I first defined which parameters I wanted to optimize and within what scope. Then, I followed the syntax for using RandomizedSearchCV:
# Random search training: fitting model
from sklearn.model_selection import RandomizedSearchCV
# create model
rf = RandomForestClassifier()
rf_random = RandomizedSearchCV(
estimator = rf,
param_distributions = random_grid,
n_iter = 100, # num combinations to search
cv = 3, # num of folds to use for cross val
verbose = 2,
random_state = 10,
n_jobs = -1,
refit = True)
# fit model
search = rf.fit(x_train, y_train)
# get best parameters
search.get_params(deep=True)
The output was a list of each parameter and its optimized value. Here, I added those hyperparameters to a random forest model and fit a new model.
# Random search training: fitting model
hyper_rf = RandomForestClassifier(
bootstrap = True,
ccp_alpha = 0.0,
class_weight= None,
criterion = 'gini',
max_features = 'auto',
min_impurity_decrease = 0.0,
min_samples_leaf = 1,
min_samples_split = 2,
min_weight_fraction_leaf = 0.0,
n_estimators = 100,
oob_score = False,
random_state = 10,
verbose = 0,
warm_start = False
)
hyper_rf.fit(x_train, y_train)
The result was a lower accuracy by 0.01%.
- only using accuracy as a performance metric instead of precision or F1 score
- not double-checking random forest tuning even though it was low accuracy
- not waiting for GridSearchCV to run on decision tree model (took >2hrs)
Phase 2 indicates the process of creating individual models for chert and obsidian datasets because of the missing columns/features: Transparency, Angularity, and Curvature. Jose Cruz and I split up the work for this. He developed a decision tree model for the obsidian data which included two datasets/experiments. I developed a decision tree model for chert data, which was contained in three datasets.
- loaded data: loaded three excel files of measurements of chert microdebitage samples that represented three unique and different stages in the knapping process into separate dataframes.
- checked for missing data: there was no missing data in any of the dataframes.
- assigned an integer for each tool stage: this is the column we will predict on
# add target column, stage, to each dataframe
chert_hh_1['Stage'] = 1
chert_hh_2['Stage'] = 2
chert_sh['Stage'] = 3
- merged data for tool stages into one large dataframe
# join dataframes
dfs = [chert_hh_1, chert_hh_2, chert_sh]
chert_df = pd.concat(dfs)
- cleaned the merged dataframe:
# remove unecessary columns + row 0
chert_df.drop(columns = ['Id',
'Filter0',
'Filter1',
'Filter2',
'Filter3',
'Filter4',
'Filter5',
'Filter6',
'hash',
'Img Id'], inplace = True)
chert_df.drop(0, inplace = True)
- filled missing data
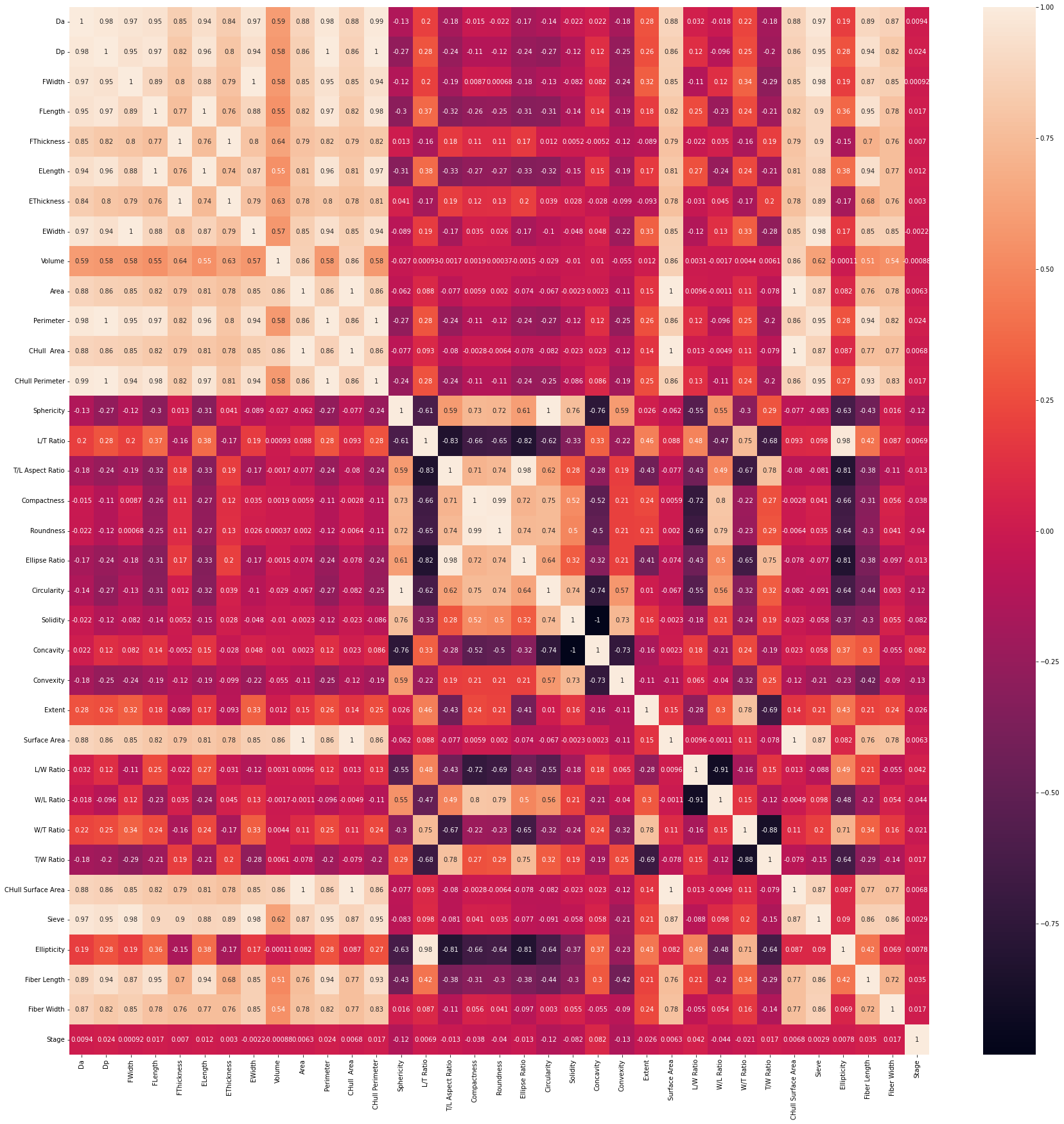
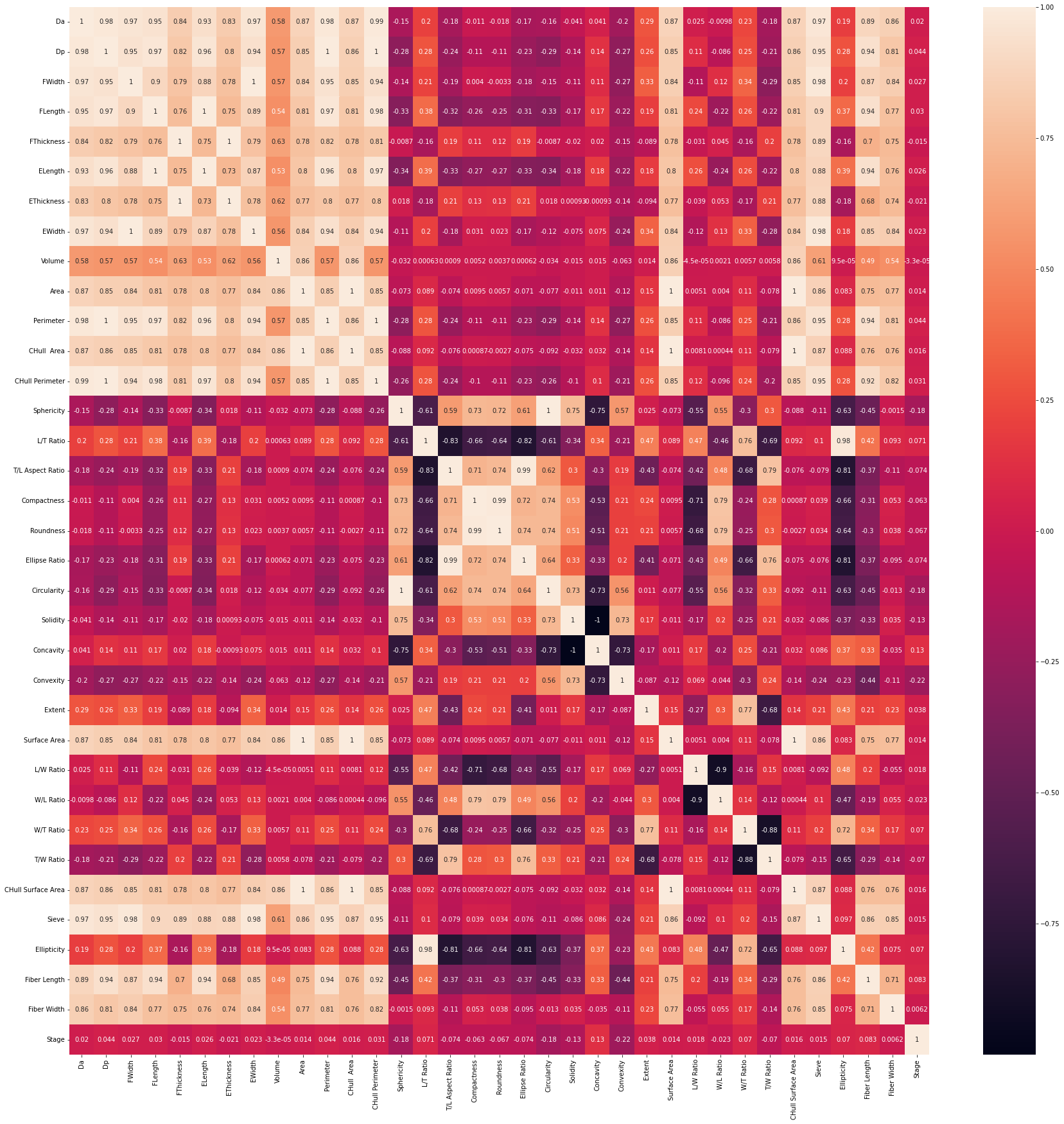
- visualized correlation between columns:
# show heatmap
import seaborn as sns
import matplotlib.pyplot as plt
plt.figure(figsize = (30,30))
sns.heatmap(chert_df.corr(), annot = True)
At this stage in our research, I was able to run an untuned decision tree model that resulted in an accuracy of 43.7%. I attempted to tune the model using GridSearchCV overnight but it resulted in a runtime error after 5 hours. I also ran an untuned random forest model which resulted in a 53.8% accuracy. The syntax did not change from Phase 1.
Next Steps involve experimenting with Convolutional Neural Networks while we wait for the chert samples to be processed again to add those three columns' data. After, we may try decision tree and random forest modeling again. Thank you!