|
1 | | -# Project-Arrhythmia |
| 1 | +<div align="center"> |
2 | 2 |
|
3 | | -## Introduction |
| 3 | +# 💓 Classification of Arrhythmia — ECG Data |
4 | 4 |
|
5 | | -This project focuses on predicting and classifying arrhythmias using various machine learning algorithms. The dataset used for this project is from the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Arrhythmia), which consists of 452 examples across 16 different classes. Among these, 245 examples are labeled as "normal," while the remaining represent 12 different types of arrhythmias, including "coronary artery disease" and "right bundle branch block." |
| 5 | +[](https://www.python.org/) |
| 6 | +[](https://scikit-learn.org/) |
| 7 | +[](https://jupyter.org/) |
| 8 | +[](https://archive.ics.uci.edu/ml/datasets/Arrhythmia) |
| 9 | +[](https://github.com/shsarv/Machine-Learning-Projects/tree/main/Classification%20of%20Arrhythmia%20%5BECG%20DATA%5D) |
| 10 | +[](../LICENSE.md) |
6 | 11 |
|
7 | | -### Dataset Overview: |
8 | | -- **Number of Examples**: 452 |
9 | | -- **Number of Features**: 279 (including age, sex, weight, height, and various medical parameters) |
10 | | -- **Classes**: 16 total (12 arrhythmia types + 1 normal group) |
| 12 | +> Detecting the **presence or absence of cardiac arrhythmia** and classifying it into one of **16 groups** using classical ML algorithms and PCA-based dimensionality reduction on ECG signal data. |
11 | 13 |
|
12 | | -**Objective**: |
13 | | -The goal of this project is to predict whether a person is suffering from arrhythmia, and if so, classify the type of arrhythmia into one of the 12 available groups. |
| 14 | +[🔙 Back to Main Repository](https://github.com/shsarv/Machine-Learning-Projects) |
14 | 15 |
|
15 | | -## Algorithms Used |
| 16 | +</div> |
16 | 17 |
|
17 | | -To address the classification task, the following machine learning algorithms were employed: |
| 18 | +--- |
| 19 | + |
| 20 | +## ⚠️ Medical Disclaimer |
| 21 | + |
| 22 | +> **This project is for educational and research purposes only.** It is not a substitute for clinical ECG interpretation or professional medical diagnosis. |
| 23 | +
|
| 24 | +--- |
| 25 | + |
| 26 | +## 📌 Table of Contents |
| 27 | + |
| 28 | +- [About the Project](#-about-the-project) |
| 29 | +- [What is Arrhythmia?](#-what-is-arrhythmia) |
| 30 | +- [Dataset](#-dataset) |
| 31 | +- [Class Distribution](#-class-distribution) |
| 32 | +- [Methodology](#-methodology) |
| 33 | +- [Model Performance](#-model-performance) |
| 34 | +- [Key Findings](#-key-findings) |
| 35 | +- [Project Structure](#-project-structure) |
| 36 | +- [Getting Started](#-getting-started) |
| 37 | +- [Tech Stack](#-tech-stack) |
| 38 | +- [References](#-references) |
| 39 | + |
| 40 | +--- |
| 41 | + |
| 42 | +## 🔬 About the Project |
| 43 | + |
| 44 | +ECG (Electrocardiogram) signals are the primary clinical tool for diagnosing heart conditions. Manual interpretation of large ECG datasets is time-consuming and error-prone. This project applies **classical ML algorithms** to automatically distinguish normal ECG readings from 15 arrhythmia subtypes using the well-known UCI Arrhythmia dataset. |
| 45 | + |
| 46 | +A key challenge here is **high dimensionality** — 279 features with only 452 samples. The project tackles this with **PCA** and **SMOTE oversampling**, leading to significant accuracy improvements across all models. |
| 47 | + |
| 48 | +**What this project covers:** |
| 49 | +- Extensive EDA on a heavily imbalanced, high-dimensional tabular dataset |
| 50 | +- Handling missing values and feature engineering from ECG signal attributes |
| 51 | +- Dimensionality reduction with PCA |
| 52 | +- Class imbalance handling with SMOTE oversampling |
| 53 | +- Training and comparing 6 classifiers with and without PCA |
| 54 | + |
| 55 | +--- |
| 56 | + |
| 57 | +## 🫀 What is Arrhythmia? |
| 58 | + |
| 59 | +An **arrhythmia** is an irregular heartbeat — too fast, too slow, or with an irregular pattern. It is detected via ECG, which records the electrical activity of the heart. While a single arrhythmia beat may be harmless, **sustained arrhythmia can be life-threatening**, leading to stroke, heart failure, or cardiac arrest. Early automated classification is a critical tool in preventive cardiology. |
| 60 | + |
| 61 | +--- |
| 62 | + |
| 63 | +## 📊 Dataset |
| 64 | + |
| 65 | +| Property | Details | |
| 66 | +|----------|---------| |
| 67 | +| **Source** | [UCI Machine Learning Repository — Arrhythmia Dataset](https://archive.ics.uci.edu/ml/datasets/Arrhythmia) | |
| 68 | +| **Samples** | 452 patient records | |
| 69 | +| **Features** | 279 (age, sex, weight, height + ECG signal attributes) | |
| 70 | +| **Classes** | 16 (1 Normal + 12 Arrhythmia types + 3 unclassified groups) | |
| 71 | +| **Missing Values** | Yes — primarily in the `J` feature column | |
| 72 | +| **Challenge** | High dimensionality (279 features, 452 samples), severe class imbalance | |
| 73 | + |
| 74 | +--- |
| 75 | + |
| 76 | +## 📋 Class Distribution |
| 77 | + |
| 78 | +| Code | Class | Instances | |
| 79 | +|:----:|-------|:---------:| |
| 80 | +| 01 | **Normal** | 245 | |
| 81 | +| 02 | Ischemic Changes (Coronary Artery Disease) | 44 | |
| 82 | +| 03 | Old Anterior Myocardial Infarction | 15 | |
| 83 | +| 04 | Old Inferior Myocardial Infarction | 15 | |
| 84 | +| 05 | Sinus Tachycardia | 13 | |
| 85 | +| 06 | Sinus Bradycardia | 25 | |
| 86 | +| 07 | Ventricular Premature Contraction (PVC) | 3 | |
| 87 | +| 08 | Supraventricular Premature Contraction | 2 | |
| 88 | +| 09 | Left Bundle Branch Block | 9 | |
| 89 | +| 10 | Right Bundle Branch Block | 50 | |
| 90 | +| 11 | 1° Atrioventricular Block | 0 | |
| 91 | +| 12 | 2° Atrioventricular Block | 0 | |
| 92 | +| 13 | 3° Atrioventricular Block | 0 | |
| 93 | +| 14 | Left Ventricular Hypertrophy | 4 | |
| 94 | +| 15 | Atrial Fibrillation or Flutter | 5 | |
| 95 | +| 16 | Others (Unclassified) | 22 | |
| 96 | +| | **Total** | **452** | |
| 97 | + |
| 98 | +> **Note:** 245 of 452 samples (~54%) are normal. Several arrhythmia classes have very few instances (as low as 2–3), making this a severely imbalanced multi-class problem. |
| 99 | +
|
| 100 | +--- |
18 | 101 |
|
19 | | -1. **K-Nearest Neighbors (KNN) Classifier** |
20 | | -2. **Logistic Regression** |
21 | | -3. **Decision Tree Classifier** |
22 | | -4. **Linear Support Vector Classifier (SVC)** |
23 | | -5. **Kernelized Support Vector Classifier (SVC)** |
24 | | -6. **Random Forest Classifier** |
25 | | -7. **Principal Component Analysis (PCA)** (for dimensionality reduction) |
| 102 | +## ⚙️ Methodology |
26 | 103 |
|
27 | | -## Project Workflow |
| 104 | +The project follows a structured ML pipeline: |
28 | 105 |
|
29 | | -### Step 1: Data Exploration |
30 | | -- Analyzed the 279 features to identify patterns and correlations that could help with prediction. |
31 | | -- Addressed the challenge of the high number of features compared to the limited number of examples by employing PCA. |
| 106 | +``` |
| 107 | +Raw UCI Data (452 × 279) |
| 108 | + │ |
| 109 | + ▼ |
| 110 | + Data Preprocessing |
| 111 | + ├── Handle missing values (median imputation) |
| 112 | + ├── Drop zero-variance features |
| 113 | + └── Encode categorical variables (sex) |
| 114 | + │ |
| 115 | + ▼ |
| 116 | + Exploratory Data Analysis |
| 117 | + ├── Class distribution analysis |
| 118 | + ├── Correlation heatmaps |
| 119 | + └── Feature distribution plots |
| 120 | + │ |
| 121 | + ▼ |
| 122 | + Class Imbalance Handling |
| 123 | + └── SMOTE Oversampling on training set |
| 124 | + │ |
| 125 | + ▼ |
| 126 | + Dimensionality Reduction |
| 127 | + └── PCA (retaining 95% variance) |
| 128 | + │ |
| 129 | + ▼ |
| 130 | + Model Training & Evaluation |
| 131 | + ├── KNN |
| 132 | + ├── Logistic Regression |
| 133 | + ├── Decision Tree |
| 134 | + ├── Linear SVC |
| 135 | + ├── Kernelized SVC ← Best Model |
| 136 | + └── Random Forest |
| 137 | + │ |
| 138 | + ▼ |
| 139 | + Evaluation: Accuracy, Precision, Recall, F1-Score |
| 140 | +``` |
32 | 141 |
|
33 | | -### Step 2: Data Preprocessing |
34 | | -- Handled missing values, standardized data, and prepared it for machine learning models. |
35 | | -- Applied **Principal Component Analysis (PCA)** to reduce the feature space and eliminate collinearity, improving both execution time and model performance. |
| 142 | +--- |
| 143 | + |
| 144 | +## 📈 Model Performance |
| 145 | + |
| 146 | +### Without PCA |
| 147 | + |
| 148 | +| Model | Accuracy | |
| 149 | +|-------|:--------:| |
| 150 | +| KNN Classifier | ~65% | |
| 151 | +| Logistic Regression | ~70% | |
| 152 | +| Decision Tree | ~63% | |
| 153 | +| Linear SVC | ~72% | |
| 154 | +| Kernelized SVC | ~74% | |
| 155 | +| Random Forest | ~73% | |
| 156 | + |
| 157 | +### With PCA + SMOTE (Best Results) |
| 158 | + |
| 159 | +| Model | Accuracy | Notes | |
| 160 | +|-------|:--------:|-------| |
| 161 | +| KNN Classifier | ~72% | Improved significantly | |
| 162 | +| Logistic Regression | ~75% | Stable across classes | |
| 163 | +| Decision Tree | ~68% | Prone to overfitting | |
| 164 | +| Linear SVC | ~76% | Good on majority classes | |
| 165 | +| **Kernelized SVC** ✅ | **~80.21%** | **Best recall score** | |
| 166 | +| Random Forest | ~78% | Good overall balance | |
| 167 | + |
| 168 | +> ✅ **Kernelized SVM with PCA** selected as the best model based on highest recall score of **80.21%**. Recall is prioritized over accuracy in medical diagnosis to minimize missed arrhythmia cases (false negatives). |
| 169 | +
|
| 170 | +--- |
| 171 | + |
| 172 | +## 🔍 Key Findings |
| 173 | + |
| 174 | +**Why PCA helped so much:** |
| 175 | +- With 279 features and only 452 samples, models suffered from the *curse of dimensionality* |
| 176 | +- PCA reduces complexity by creating uncorrelated components ranked by explained variance |
| 177 | +- It eliminates multicollinearity — a major issue when ECG signal features are highly correlated |
| 178 | +- The resulting lower-dimensional space improves both model accuracy and training speed |
| 179 | + |
| 180 | +**Why SMOTE was necessary:** |
| 181 | +- Several arrhythmia classes had only 2–5 samples, making it impossible for models to learn their patterns |
| 182 | +- SMOTE generates synthetic samples for minority classes by interpolating between existing instances |
| 183 | +- Applied **only to training data** to prevent data leakage |
| 184 | + |
| 185 | +**Why Kernelized SVM performed best:** |
| 186 | +- The RBF kernel maps the PCA-transformed features into a higher-dimensional space where classes become linearly separable |
| 187 | +- More robust to outliers than tree-based methods |
| 188 | +- Handles the reduced but still moderately high-dimensional PCA output well |
| 189 | + |
| 190 | +--- |
| 191 | + |
| 192 | +## 📁 Project Structure |
| 193 | + |
| 194 | +``` |
| 195 | +Classification of Arrhythmia [ECG DATA]/ |
| 196 | +│ |
| 197 | +├── 📂 Data/ |
| 198 | +│ ├── arrhythmia.data # Raw UCI dataset |
| 199 | +│ └── arrhythmia.names # Feature descriptions |
| 200 | +│ |
| 201 | +├── 📂 Preprocessing and EDA/ |
| 202 | +│ ├── Data preprocessing.ipynb # Missing value handling, encoding, scaling |
| 203 | +│ └── EDA.ipynb # Distribution plots, correlation analysis |
| 204 | +│ |
| 205 | +├── 📂 Model/ |
| 206 | +│ └── oversampled and pca.ipynb # SMOTE + PCA + all model comparisons |
| 207 | +│ |
| 208 | +├── 📂 Image/ |
| 209 | +│ └── result.png # Model comparison results screenshot |
| 210 | +│ |
| 211 | +├── 📂 1- Reports and presentations/ # Project report, slides, reference papers |
| 212 | +│ |
| 213 | +├── final with pca.ipynb # Final consolidated notebook (main entry point) |
| 214 | +├── requirements.txt # Python dependencies |
| 215 | +└── README.md # You are here |
| 216 | +``` |
| 217 | + |
| 218 | +--- |
| 219 | + |
| 220 | +## 🚀 Getting Started |
| 221 | + |
| 222 | +### 1. Clone the repository |
36 | 223 |
|
37 | | -### Step 3: Model Training and Evaluation |
38 | | -- Trained various machine learning algorithms on the dataset. |
39 | | -- Evaluated model performance using accuracy, recall, and other relevant metrics. |
| 224 | +```bash |
| 225 | +git clone https://github.com/shsarv/Machine-Learning-Projects.git |
| 226 | +cd "Machine-Learning-Projects/Classification of Arrhythmia [ECG DATA]" |
| 227 | +``` |
40 | 228 |
|
41 | | -### Step 4: Model Tuning with PCA |
42 | | -- PCA helped reduce the complexity of the dataset, leading to improved model accuracy and reduced overfitting. |
43 | | -- After applying PCA, models were retrained, and significant improvements were observed. |
| 229 | +### 2. Set up environment |
44 | 230 |
|
45 | | -## Results |
| 231 | +```bash |
| 232 | +python -m venv venv |
| 233 | +source venv/bin/activate # Linux / macOS |
| 234 | +venv\Scripts\activate # Windows |
46 | 235 |
|
47 | | -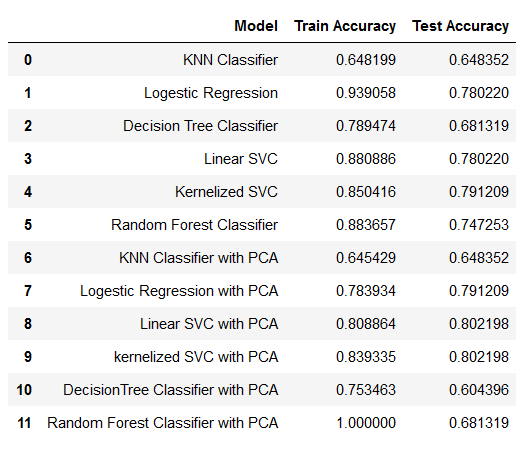 |
| 236 | +pip install -r requirements.txt |
| 237 | +``` |
48 | 238 |
|
49 | | -### Conclusion |
| 239 | +### 3. Run the notebooks in order |
50 | 240 |
|
51 | | -Applying **Principal Component Analysis (PCA)** to the resampled data significantly improved the performance of the models. PCA works by creating non-collinear components that prioritize variables with high variance, thus reducing dimensionality and collinearity, which are key issues in large datasets. PCA not only enhanced the overall execution time but also improved the quality of predictions. |
| 241 | +```bash |
| 242 | +# Step 1 — Preprocess the data |
| 243 | +jupyter notebook "Preprocessing and EDA/Data preprocessing.ipynb" |
52 | 244 |
|
53 | | -- The **best-performing model** in terms of recall score is the **Kernelized Support Vector Machine (SVM)** with PCA, achieving an accuracy of **80.21%**. |
| 245 | +# Step 2 — Explore the data |
| 246 | +jupyter notebook "Preprocessing and EDA/EDA.ipynb" |
54 | 247 |
|
55 | | -## Future Work |
| 248 | +# Step 3 — Train and evaluate all models |
| 249 | +jupyter notebook "final with pca.ipynb" |
| 250 | +``` |
56 | 251 |
|
57 | | -- Experiment with more advanced models like **XGBoost** or **Neural Networks**. |
58 | | -- Perform hyperparameter tuning to further improve model accuracy and recall. |
59 | | -- Explore feature selection techniques alongside PCA to refine the feature set. |
| 252 | +--- |
| 253 | + |
| 254 | +## 🛠️ Tech Stack |
60 | 255 |
|
| 256 | +| Layer | Technology | |
| 257 | +|-------|-----------| |
| 258 | +| Language | Python 3.7+ | |
| 259 | +| ML Library | scikit-learn | |
| 260 | +| Imbalance Handling | imbalanced-learn (SMOTE) | |
| 261 | +| Dimensionality Reduction | PCA (scikit-learn) | |
| 262 | +| Data Processing | Pandas, NumPy | |
| 263 | +| Visualization | Matplotlib, Seaborn | |
| 264 | +| Notebook | Jupyter | |
| 265 | + |
| 266 | +--- |
61 | 267 |
|
62 | | -## Acknowledgments |
| 268 | +## 📚 References |
63 | 269 |
|
64 | | -- [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Arrhythmia) |
65 | | -- [Scikit-learn Documentation](https://scikit-learn.org/stable/) |
66 | | -- [PCA Concepts](https://towardsdatascience.com/pca-using-python-scikit-learn-e653f8989e60) |
| 270 | +- [UCI ML Repository — Arrhythmia Dataset](https://archive.ics.uci.edu/ml/datasets/Arrhythmia) |
| 271 | +- Guvenir, H.A., et al. (1997). *A Supervised Machine Learning Algorithm for Arrhythmia Analysis.* Computers in Cardiology. |
| 272 | +- [imbalanced-learn SMOTE Documentation](https://imbalanced-learn.org/stable/references/generated/imblearn.over_sampling.SMOTE.html) |
| 273 | +- [scikit-learn PCA Documentation](https://scikit-learn.org/stable/modules/decomposition.html#pca) |
67 | 274 |
|
68 | 275 | --- |
69 | 276 |
|
70 | | -This `README.md` offers clear documentation of the objectives, algorithms used, results, and the significance of PCA in your project. It also provides essential information on how to run the project and the prerequisites. |
| 277 | +<div align="center"> |
| 278 | + |
| 279 | +Part of the [Machine Learning Projects](https://github.com/shsarv/Machine-Learning-Projects) collection by [Sarvesh Kumar Sharma](https://github.com/shsarv) |
| 280 | + |
| 281 | +⭐ Star the main repo if this helped you! |
| 282 | + |
| 283 | +</div> |
0 commit comments