I want to batch multiple small jobs together to reduce cluster queue wait times. My cluster administrators don't want jobs that finish really quickly being queued due to strain on the slurm scheduler. In this example, each job runs quickly (<1 minute) . Queuing thousands of these jobs on a busy cluster sucks. Each job might take several minutes or longer to start, then it is done in less than a minute. I'd prefer when getting a reservation for more like ~20-100 of these jobs to run in serial. But I have other rules that take longer, so I can't resort to --cores=1 in the snakemake invocation.
Example workflow:
rule trim_primers:
input:
forward = "input/{run_pe}-{sample_pe}_R1.fastq.gz",
reverse = "input/{run_pe}-{sample_pe}_R2.fastq.gz"
output:
forward_trimmed="output/v3v4_trimmed/{run_pe}-{sample_pe}_R1.trimmed.fastq.gz",
reverse_trimmed="output/v3v4_trimmed/{run_pe}-{sample_pe}_R2.trimmed.fastq.gz"
resources:
cpus_per_task=1,
mem_mb=1000,
runtime="30m",
slurm_partition="short"
shell:
...
rule map_reads:
input:
forward = "output/v3v4_trimmed/{run_pe}-{sample_pe}_R1.trimmed.fastq.gz",
reverse = "output/v3v4_trimmed/{run_pe}-{sample_pe}_R2.trimmed.fastq.gz"
output:
forward_trimmed="output/mapped/{run_pe}-{sample_pe}_mapped.bam",
reverse_trimmed="output/mapped/{run_pe}-{sample_pe}_mapped.bam"
resources:
cpus_per_task=8,
mem_mb=16000,
runtime="8h",
slurm_partition="short"
shell:
...When using Snakemake's grouping feature:
snakemake \
--groups trim_primers=group_trim_primers \
--group-components group_trim_primers=20
Snakemake requests a node with 20 CPUs and 20GB RAM to run all jobs within the group in parallel.
Current behavior (from docs):
Snakemake will request resources for groups by summing across jobs that can be run in parallel
However, I'd like the following behavior:
Desired behavior:
Run batches of 20 samples sequentially on the same node using only 1 CPU and 1GB RAM total. Ideally I wouldn't have to change the time either and could configure the group to have a time limit altogether.
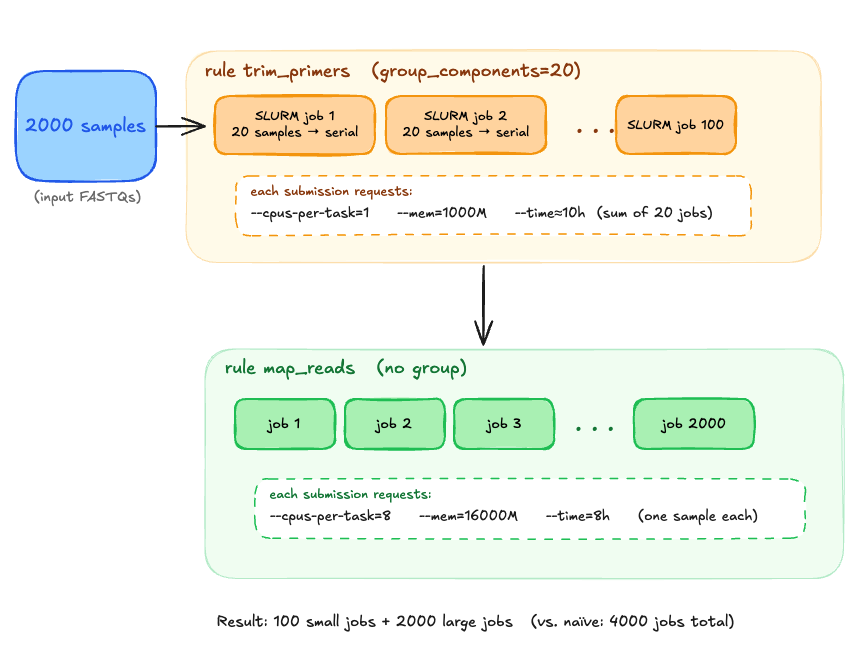
Is there a way to configure Snakemake groups to run jobs sequentially rather than in parallel, without resorting to a wrapper rule that does the batching in python? If not, is this a Snakemake feature request or one for slurm executor plugin?
I want to batch multiple small jobs together to reduce cluster queue wait times. My cluster administrators don't want jobs that finish really quickly being queued due to strain on the slurm scheduler. In this example, each job runs quickly (<1 minute) . Queuing thousands of these jobs on a busy cluster sucks. Each job might take several minutes or longer to start, then it is done in less than a minute. I'd prefer when getting a reservation for more like ~20-100 of these jobs to run in serial. But I have other rules that take longer, so I can't resort to
--cores=1in the snakemake invocation.Example workflow:
When using Snakemake's grouping feature:
Snakemake requests a node with 20 CPUs and 20GB RAM to run all jobs within the group in parallel.
Current behavior (from docs):
However, I'd like the following behavior:
Desired behavior:
Run batches of 20 samples sequentially on the same node using only 1 CPU and 1GB RAM total. Ideally I wouldn't have to change the time either and could configure the group to have a time limit altogether.
Is there a way to configure Snakemake groups to run jobs sequentially rather than in parallel, without resorting to a wrapper rule that does the batching in python? If not, is this a Snakemake feature request or one for slurm executor plugin?